Bio-marker Sourcing of BatsSpecies Distribution ModelRe-purposed Data for phylogeography |
Bio-marker Sourcing of Bats
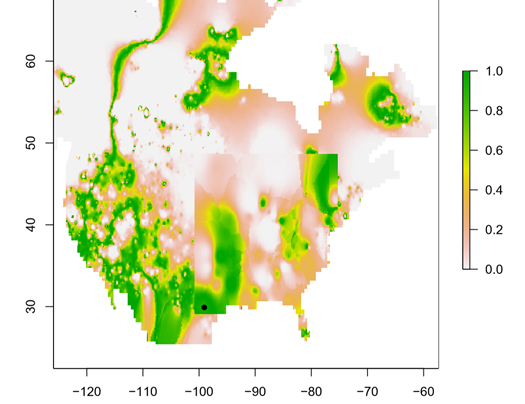
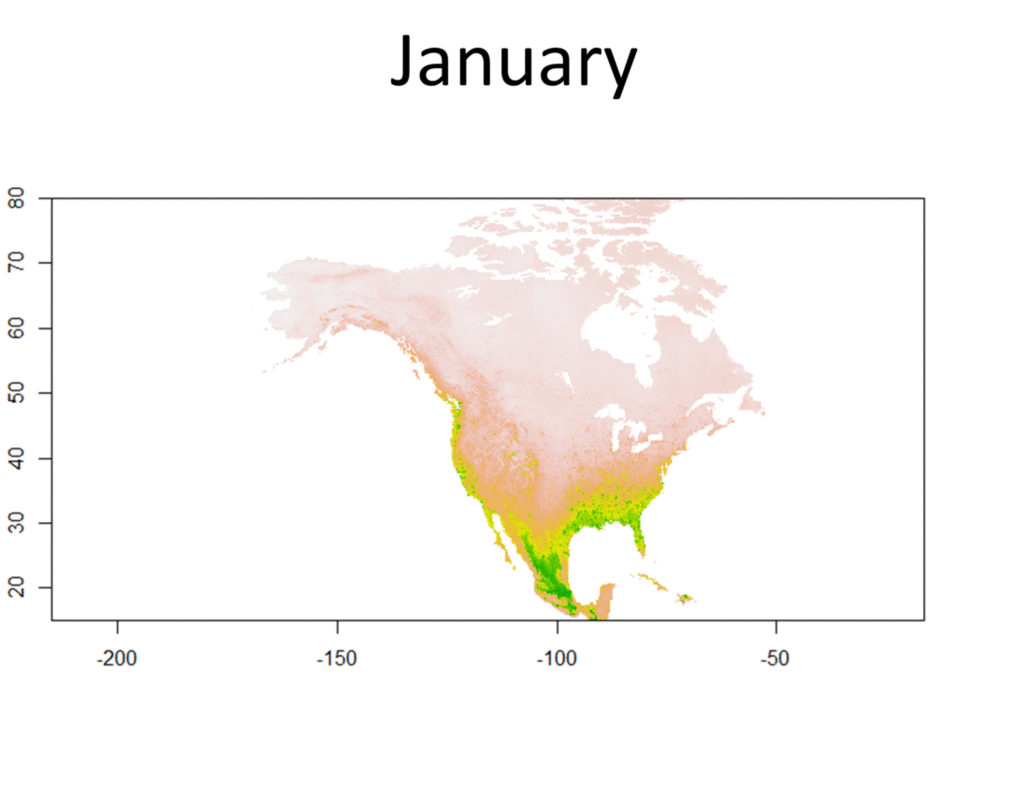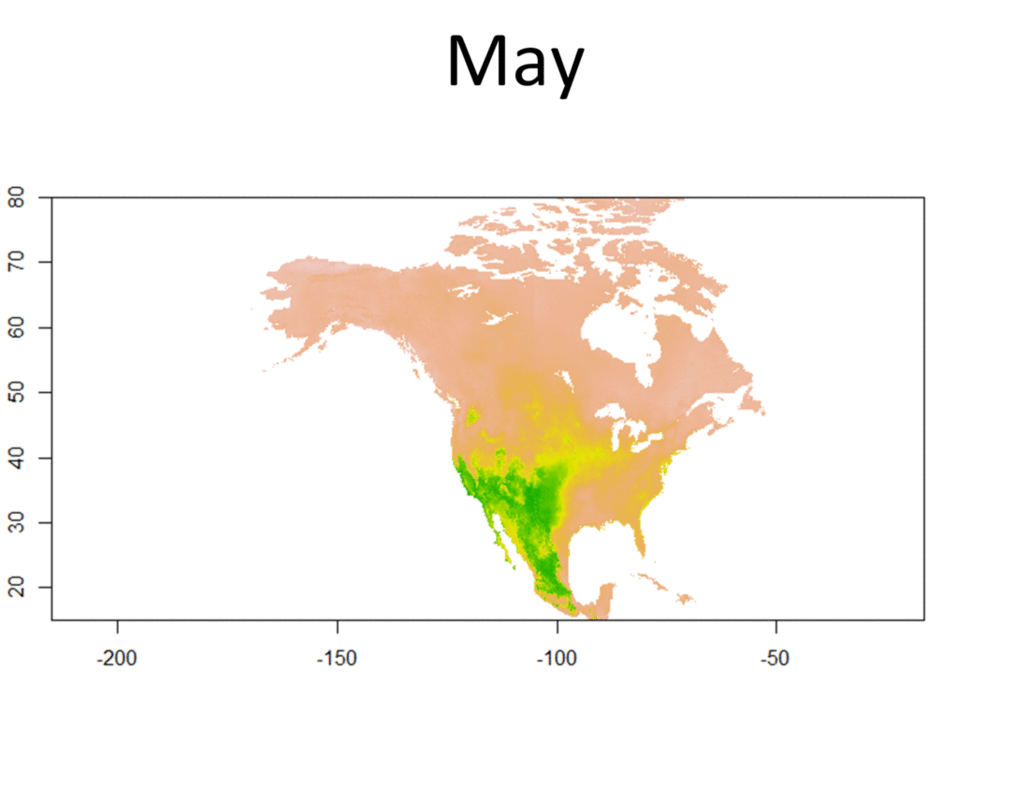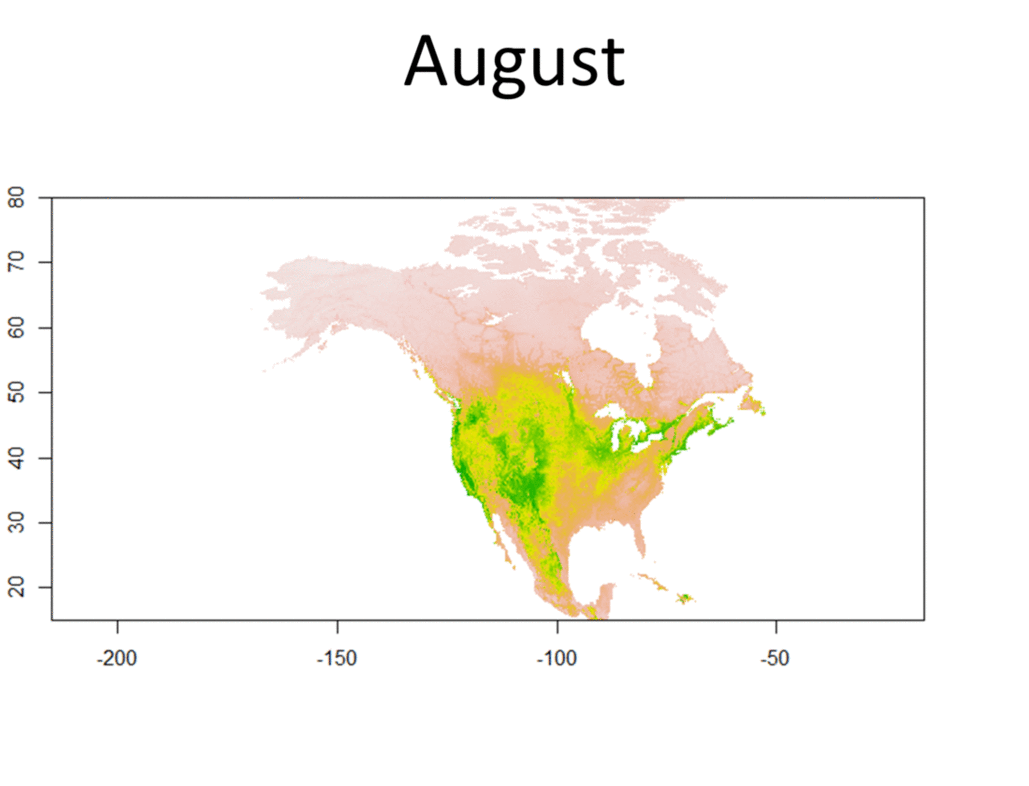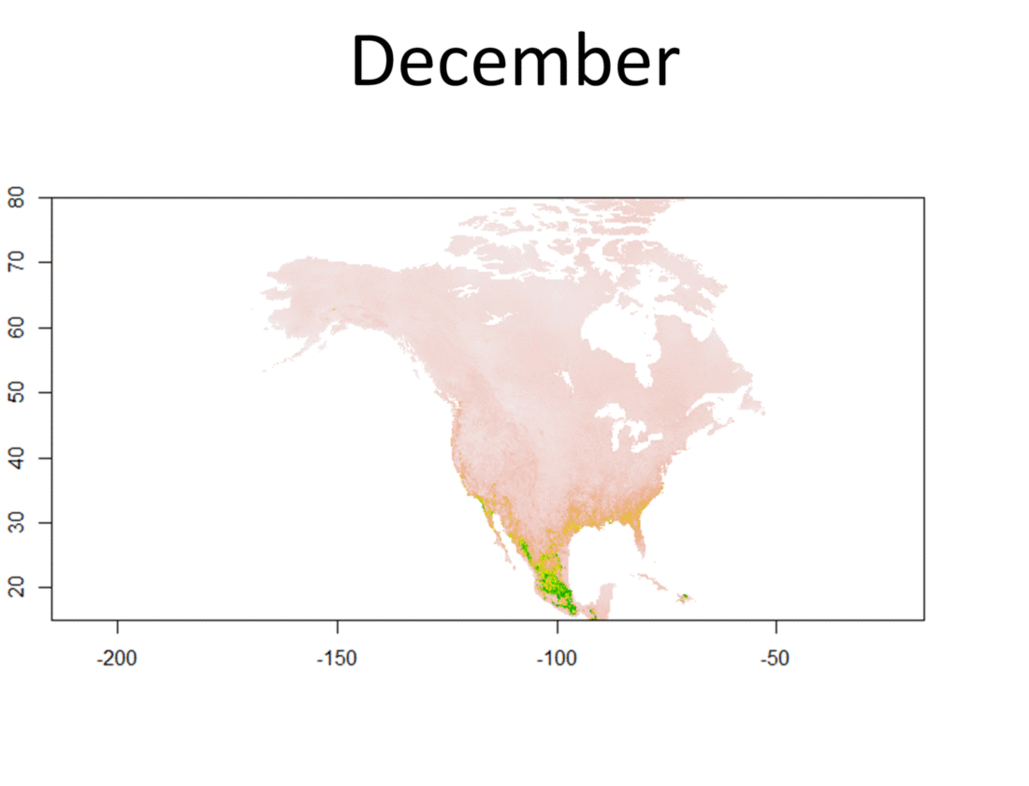
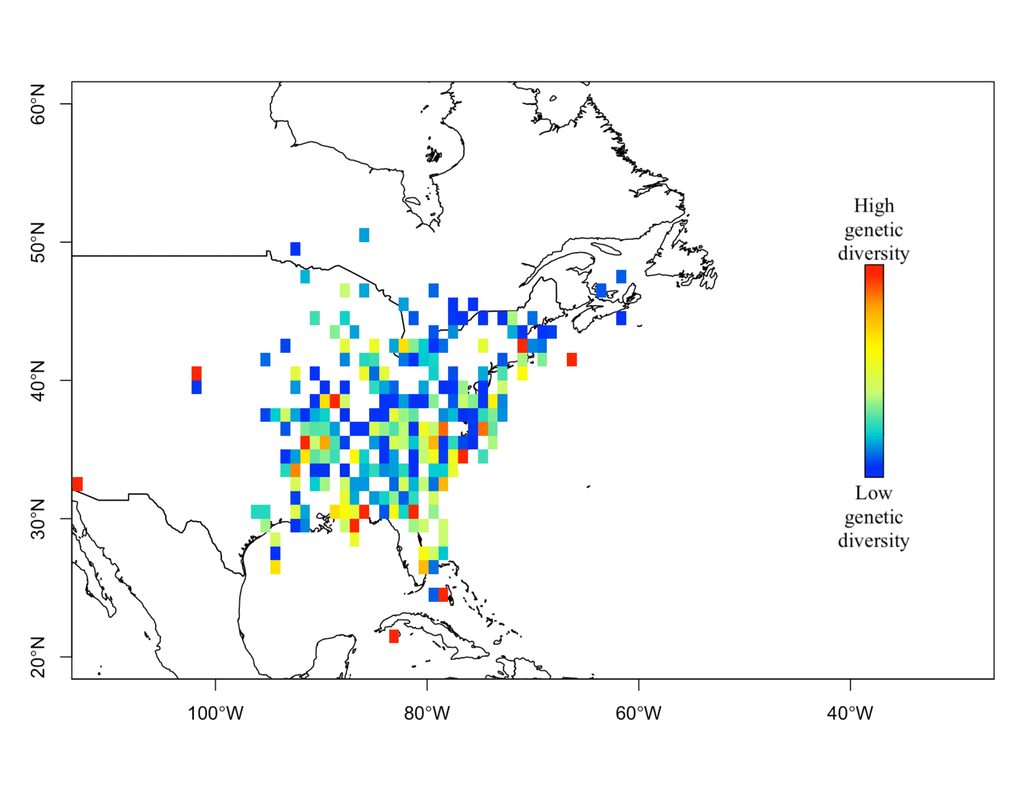
My research focuses on bats killed at wind turbines. My dissertation involves attempting to determine the origin of bats killed at wind farms. Three types of data are used for this project, trace element, isotope, and genetic data. Isotopes have been used greatly in the past to track the movements of migratory animals, and genetics have provided some insight as well. Meanwhile, only recently have studies been able to use trace element information from biological samples to track the movements of wildlife. Species Distribution and Re-purposed Data Another focus of my research is the use of re-purposed data to answer new and interesting questions. This has primarily occurred in three areas. The first, and relating to the bio-marker research, is the use of Hoary, Silver-haired, and Eastern Red Bat occurances from GBIF (https://www.gbif.org/) to create seasonal species distribution models to better understand how these species move through the environment. Second, I am assisting in a project led by Danielle Parsons, with another colleague Drew Duckett, using GBIF and GenBank data to better understand the cryptic diversity present in mammals. Lastly, the use of data from GenBank and GBIF to understand the phylogeographic history of the Southeast US. eDNA and Invasive Species Previous research for my master's degree at Central Michigan University was conducted on invasive grass carp in the Western Basin of Lake Erie. This was primarily concerned with using eDNA to help determine their locations and using blood samples from captured fish to determine their ploidy and therefore their reproductive viability in the Western Basin. |
|
|
|
|
|
|